A recent publication in Journal of Experimental Medicine by the Sallusto group explored the nature of the human CD4+ T cell immunodominance to influenza H1 hemagglutinin and demonstrate that the naïve repertoire is very broad, while the memory repertoire becomes selected toward peptides with a high likelihood to be processed and presented by dendritic cells.
Influenza viruses are responsible every year of significant economic loss as well as severe morbidity and mortality, especially in children <5 yr old and adults >65. Moreover, the rapidly evolving nature of influenza viruses poses risks of serious epidemics or pandemics. CD4+ T cells are key components of protective immune responses to viruses, being necessary for the elicitation of antibody and cytotoxic T cell responses. However, the mechanisms that determine immunogenicity and the selection of immunodominant epitopes within complex protein antigens remain elusive.
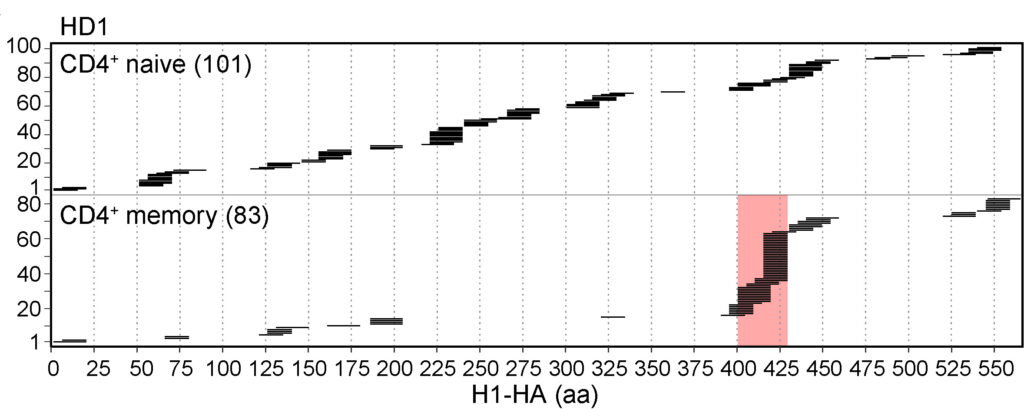
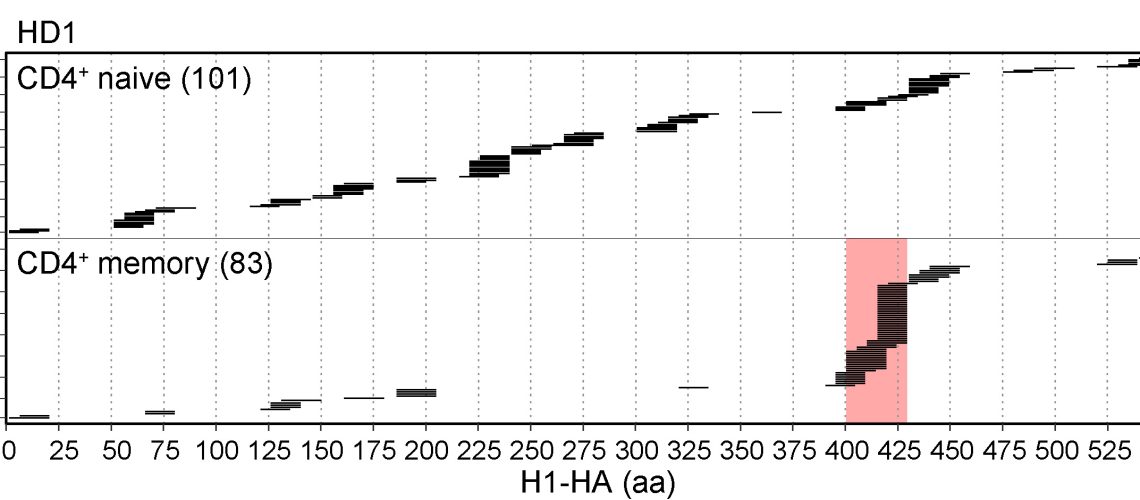
By analyzing antigen presentation and CD4+ T cell recognition of influenza pandemic H1 hemagglutinin (H1-HA), the international team of researchers from the Institute for Research in Biomedicine (Università della Svizzera italiana) and the Institute of Microbiology (ETH Zürich) dissected the impact of antigen processing on T cell clonal selection and immunodominance. In particular, they found that naive CD4+ T cells have a very broad repertoire, being able to recognize naturally processed as well as cryptic peptides spanning the whole H1-HA sequence. In contrast, memory Th cells were primarily directed against just a few immunodominant peptides that were readily detected by mass spectrometry-based MHC-II peptidomics and predicted by structural accessibility analysis.
Importantly, this work proposes methods to improve in silico predictors of CD4+ T cell responses to viral infectious diseases. A better understanding of the rules governing immunodominance and T cell clonal selection has indeed a profound impact in regard of vaccine design and preparedness for future epidemics and pandemics.
This work, published in the renowned “Journal of Experimental Medicine ”, was realized in collaboration with researchers from the Tulane University (New Orleans, LA, USA), from the Swiss Institute of Bioinformatics (Lausanne) and from the Surrey Clinical Research Centre (University of Surrey, Guildford, UK). The study was financially supported by the European Research Council, the Swiss National Science Foundation, and the European Union.
Article:
Deciphering and predicting CD4+ T cell immunodominance of influenza virus hemagglutinin
A. Cassotta, P. Paparoditis, R. Geiger, R. R. Mettu, S. J. Landry, A. Donati, M. Benevento, M. Foglierini, D. J. M. Lewis, A. Lanzavecchia, F. Sallusto
in J Exp Med (2020) vol. 217